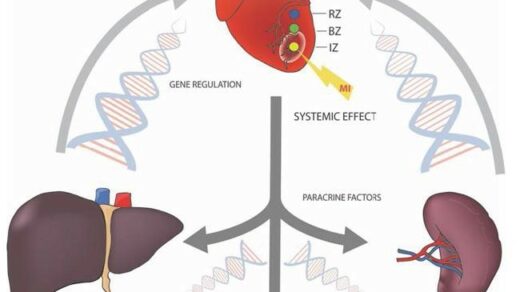
In this trending new study, researchers used CRISPR-based genome-wide screens to identify genetic determinants of PARP10-mediated cellular survival.
The Trending With Impact series highlights Oncotarget publications attracting higher visibility among readers around the world online, in the news, and on social media—beyond normal readership levels. Look for future science news about the latest trending publications here, and at Oncotarget.com.
—
Genetic interactions involved in the survival of cancer cells are potential therapeutic targets in personalized cancer therapy. “Synthetic lethal” is a type of genetic interaction where the knockout of one gene can cause cell death but only in the presence of another dependent gene. Cancer researchers view synthetic lethality screening as a powerful tool in precision medicine.
“Identifying genetic susceptibilities based on PARP10 expression levels is thus potentially relevant for finding new targets for precision oncology.”
Poly-ADP-ribose polymerase 10, or PARP10, is a nuclear protein that is overexpressed in multiple cancers. Genetic susceptibilities based on PARP10 expression levels in an individual may be potential targets for personalized cancer therapy. In a new study, researchers Jude B. Khatib, Emily M. Schleicher, Lindsey M. Jackson, Ashna Dhoonmoon, George-Lucian Moldovan, and Claudia M. Nicolae, from the Department of Biochemistry and Molecular Biology at Penn State College of Medicine, used CRISPR-based, genome-wide genetic screens to identify potential synthetic lethality interactions with PARP10-overexpressing and -knockout cancer cells. On September 28, 2022, their research paper was published in Oncotarget and entitled, “Complementary CRISPR genome-wide genetic screens in PARP10-knockout and overexpressing cells identify synthetic interactions for PARP10-mediated cellular survival.”
“Here, we employed complementary CRISPR loss-of-function genome-wide screening to identify genes required for proliferation of PARP10-overexpressing and PARP10-knockout cells.”
The Study
To identify potential synthetic lethal targets, the researchers conducted a CRISPR-based, genome-wide genetic screen of both PARP10-overexpressing and PARP10-knockout tumorigenic and non-tumorigenic breast cells. The screen looked for genes that were required for cell proliferation in the presence of PARP10 overexpression or PARP10 knockout.
“Here, we performed a series of CRISPR genome-wide loss-of-function screens in isogenic control and PARP10-overexpressing or PARP10-knockout cell lines, to identify genetic determinants of PARP10-mediated cellular survival.”
In the PARP10 overexpressing cells, the top results from their CRISPR screen were validated with biological pathway enrichment analyses, using both KEGG and Gene Ontology databases. A functional interaction between ATM and PARP10 expression was found. ATM promoted cell proliferation in PARP10-overexpressing cells.
In the genome-wide CRISPR knockout screens, genes required for the viability of PARP10-knockout cells were identified. In the PARP10 knockout cells, the top results from their CRISPR screen were validated with biological pathway enrichment analyses, using both KEGG and Gene Ontology databases. They identified the CDK2-Cyclin E1 complex as a genetic determinant for the proliferation of PARP10-knockout cells.
“Our work identifies a network of functionally relevant PARP10 synthetic interactions, and reveals a set of factors which can potentially be targeted in personalized cancer therapy.”
Conclusion
The researchers identified several genes that were differentially required for cell proliferation in the presence of PARP10 overexpression or knockout. Some of these genes have been previously implicated in cancer, while others were novel candidate cancer targets. The identification of these potential synthetic lethal interactions provides new insights into the role of PARP10 in cancer and may be useful for precision oncology. This study highlights the importance of using complementary CRISPR-based screens to identify potential cancer targets.
“We found that DNA repair factors, including ATM, a master regulator of the DNA damage checkpoint response, are specifically promoting the proliferation of PARP10-overexpressing cells. Moreover, we identified a role for PARP10 in regulating ATM recruitment to stressed replication forks. Finally, we found that the CDK2-cyclin E1 complex is specifically required for the proliferation of PARP10-deficient cells. Our work reveals novel PARP10 genetic interactions of functional relevance and identifies a set of factors which can potentially be targeted in personalized cancer therapy.”
Click here to read the full research paper published by Oncotarget.
ONCOTARGET VIDEOS: YouTube | LabTube | Oncotarget.com
—
Oncotarget is an open-access journal that publishes primarily oncology-focused research papers in a continuous publishing format. These papers are available at no cost to readers on Oncotarget.com. Open-access journals have the power to benefit humanity from the inside out by rapidly disseminating information that may be freely shared with researchers, colleagues, family, and friends around the world.
For media inquiries, please contact media@impactjournals.com.